A tour of the cell
Use 'Print preview' to check the number of pages and printer settings.
Print functionality varies between browsers.
Printable page generated Friday, 3 May 2024, 12:20 AM
A tour of the cell
Introduction
This course contains a blend of text and a multimedia interactive component to look at the uniformity and diversity within cells. Fundamental to understanding how cells 'work' is knowledge of the subcellular components and how they are arranged and this is tested through a series of in-text and self-assessment questions.
This OpenLearn course is an adapted extract from the Open University course S294 Cell biology.
Learning outcomes
After studying this course, you should be able to:
list the main components of cells
summarise the structure and function of the different components
outline how cell ultrastructure is related to cell function
identify cell organelles and the main cytoskeletal components in diagrams and EM micrographs and interpret simple EM images.
1 Introducing the cell
Studies into the function and appearance of cells reveal the presence of great diversity, which is particularly striking among eukaryotic cells. Even very different types of eukaryotic cells, however, exhibit many common structures and functions. This mix of uniformity and diversity is also reflected in the organisation within cells. This course looks in more detail at the interior of cells - their subcellular structure, including their 'ultrastructural' features (sometimes called 'fine structure'), which are visible using electron microscopy. Knowledge of the subcellular components of cells and how these components are arranged is fundamental to your understanding of how cells perform their functions: that is, how cells 'work'.
Schematic diagrams of typical animal and plant cells are shown in Figure 1a and b. You do not need to study this figure in detail now; it will be referred to again during the course of this course.
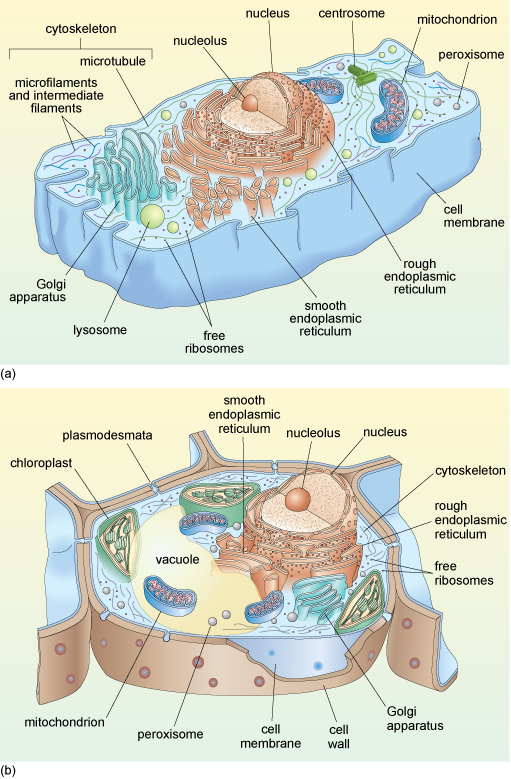
The diagrams are three-dimensional representations of the cell's internal structure. Near the centre in both cells is the nucleus, which contains a small nucleolus. The nucleus is surrounded by a double membrane, or envelope, with pores in its surface. Outside of and continuous with the nuclear envelope are the membranous sacs that make up the endoplasmic reticulum. The endoplasmic reticulum nearest the nucleus is studded with ribosomes and is described as rough; while further out from the nucleus is the smooth endoplasmic reticulum, which lacks associated ribosomes. Other membranous sacs present in both cells make up the Golgi apparatus. Also present in the cytosol are numerous free ribosomes, small spherical structures called peroxisomes and several mitochondria; the mitochondria have an outer membrane and also a highly convoluted inner membrane. Both the animal cell and the plant cell also contain cytoskeletal components - microfilaments, intermediate filaments and microtubules; and both cells are bounded by the cell membrane. Present in the animal cell' but not shown in the plant cell, are lysosomes and also a pair of small cylindrical organelles collectively called the centrosome. The plant cell differs from the animal cell in having a cell wall outside the cell membrane, which is punctuated by connections called plasmodesmata - these are channels, lined by cell membrane, that pass through the cell walls of adjacent cells. Finally, the plant cell contains several chloroplasts, oval organelles containing stacks of green, chlorophyll-containing flattened sacs called grana; the grana are linked by membranes called grana lamellae.
Before beginning your study of cell components, some of the main techniques used to study the interior organisation of cells are briefly outlined.
2 How is subcellular organisation studied?
Subcellular organisation has been studied in two main ways: by electron microscopy (EM) and by the separation (fractionation) and subsequent biochemical or molecular analysis of the different cell components.
2.1 Electron microscopy
Light microscopes cannot be used to study very small structures (i.e. those less than approximately 0.2 µm across), because there is a limit to the resolution, or 'resolving power' of such microscopes. The limiting factor that prevents small structures being discriminated using a light microscope is the wavelength of visible light.
The problem of the limited resolution of the traditional light microscope was overcome in the 1930s, by the development of electron microscopes, which use beams of electrons instead of light. Electron beams have a shorter wavelength than that of visible light, and the wavelength of a beam of electrons decreases as the velocity (speed) at which they travel increases. Electron beams accelerated at high velocity through an appropriately prepared tissue section and focused on a fluorescent screen can hence be used to resolve cellular components that are as small as 1 nm across.
There are two different types of electron microscopy. Scanning electron microscopy (SEM) is a technique used to study the surface of intact cells and tissues. The sample is coated with a thin metallic layer that deflects an electron beam onto a detector. The internal organisation of cells is studied using the second technique, transmission electron microscopy (TEM), in which a beam of electrons is passed through a very thin tissue section, allowing the analysis of cell organelles and other components.
Transmission electron microscopy
Transmission electron microscopy has been greatly refined since the first commercial electron microscope became available in 1939. TEM has allowed detailed examination of cell ultrastructure and assisted the identification and investigation of cell organelles such as the Golgi apparatus (Section 4.7), which had previously been seen only as indistinct subcellular structures using histochemical techniques and light microscopy.
As in light microscopy, samples must be fixed and processed before they can be viewed using a transmission electron microscope (Figure 2a) although the reagents used are different from those used for light microscopy. Preservation of the tissue is very important in this technique; it is crucial that the membranes of the cell organelles are preserved and not obscured by precipitates (insoluble deposits) formed during fixation. Glutaraldehyde, which fixes proteins, followed by osmium tetroxide, which fixes lipids, are the fixatives most often used. The fixed tissue is embedded in a very rigid resin, such as epoxy resin, which allows very thin sections to be cut on an ultramicrotome. 'Ultrathin' EM sections, which are used for study of cell organelles, are typically around 70 nm thick (compared with the 5-50 µm thick sections typically used for light microscopy). The image is obtained by passing electrons through the section, and focusing them on a fluorescent screen that emits visible light where electrons strike it. The interior of the microscope is under vacuum (to prevent scattering of the electron beam by air molecules) and the direction of the electron beam is controlled by magnets.
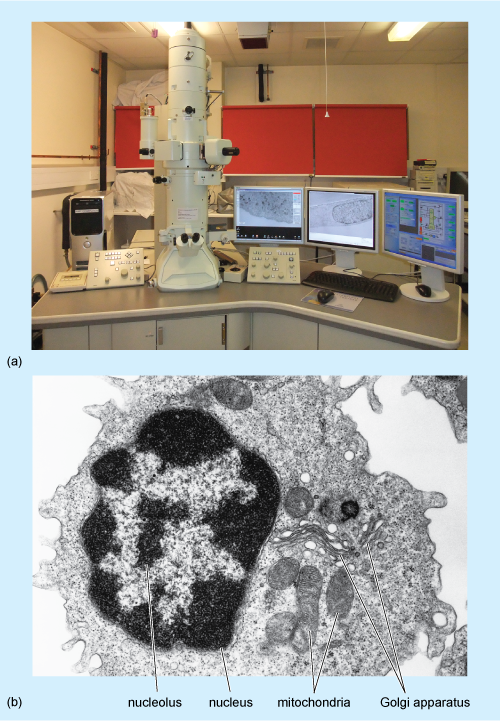
The photo in (a) shows the tall cylindrical microscope, which has a binocular eyepiece near its base. The microscope stands on the bench and the images are displayed on computer screens beside it. The electron micrograph in (b) shows a roughly spherical cell with its darkly staining (electron-dense) nucleus toward the left. Within the nucleus are several large and irregularly-shaped, pale (electron-lucent) areas, which are the nucleoli. In the cytoplasm, to the right of the nucleus, are several mitochondria which have been sectioned in different planes; the internal membranes of the organelle appear dark against the paler matrix. Also present in the cytoplasm are the fluid-filled vesicles and flattened sacs of the Golgi apparatus.
Electrons pass readily through unstained tissue sections, because the cell components are made up of small atoms such as carbon, oxygen and hydrogen. To enable cellular components such as organelles to be viewed, the sections must first be 'stained' to increase contrast. However, unlike light microscopy, which uses coloured stains that absorb light, in TEM, the stains contain heavy metals, such as lead and uranium. These are large atoms that prevent electrons passing straight through the section. Uranium binds preferentially to nucleic acids and proteins, while lead binds preferentially to lipids. So, after staining, cell components that are rich in lipids and areas where proteins and DNA are concentrated prevent the passage of electrons and so appear relatively dark, or 'electron-dense', on the viewing screen. Areas in which proteins are less concentrated, such as the cytosol of animal cells, appear pale, or 'electron-lucent'. A typical TEM image, of a white blood cell, is shown in Figure 2b.
Immunoelectron microscopy
Immunolabelling to localise specific molecules to particular cell organelles can also be applied in electron microscopy. In EM immunolabelling (often known as immunoelectron microscopy, or immuno-EM), the antibody molecules are tagged with electron-dense substances, the most effective being small gold particles, which can be observed as dark dots. Different antibodies can be tagged with particles of different sizes, allowing detection of more than one type of molecule to be performed simultaneously.
Figure 3 shows an example of how immunocytochemistry at the EM level can provide information about the localisation of molecules in an individual cell. The figure shows two cells of a bacterium, Neisseria meningitidis, labelled with an antibody that recognises a particular oligosaccharide (short chain of sugar units). The binding of this primary antibody is visualised using a secondary antibody coupled to gold particles, 15 nm in diameter, each of which can be seen as a very dark (electron-dense) dot.
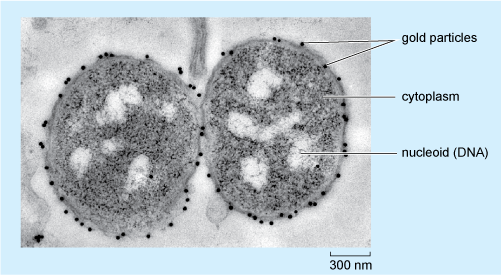
The gold particles appear as numerous black dots, all but one of which is on the outer surface of the bacteria. The pale, non-granular cell wall can be distinguished from the darker, granular cytoplasm beneath it. The nucleoid appears as a number of white patches, of a variety of sizes and shapes, within the cytoplasm.
Looking at Figure 3, in which part of the bacterium is the oligosaccharide located?
The gold particles are, with only one exception, found on the outer surface of the two bacteria; so it can be deduced that this is where the oligosaccharide is predominantly localised in N. meningitidis.
2.2 Cell fractionation
Although electron microscopy has allowed the appearance of cell organelles to be studied, elucidation of their function required a method for separating the different organelles from the remaining cell components so that their biochemical properties could be studied. This vital link between cell structure and biochemistry was made possible by developments in a technique known as cell fractionation (Box 1). This procedure is carried out in two steps: disruption of the cells to release the subcellular components, followed by centrifugation (described below) to separate the components on the basis of their mass or density.
Centrifugation
Large particles suspended in liquid will eventually settle out under the influence of the Earth's gravitational field (the strength of this force is denoted g). Small particles, however, will usually separate extremely slowly (or not at all) unless subjected to a much greater force than gravity. This can be achieved by centrifugation, in which the suspension of particles is spun round an axis at very high rotational speeds (Figure 4), creating what is commonly referred to as a centrifugal force, which causes particles to migrate away from the axis. The rate at which particles settle out (or sediment) depends on their size, shape and density (density is the mass of a given volume of a substance, i.e. density = mass/volume), the density of the liquid, and the rotational speed of centrifugation. Centrifugation at different speeds allows the separation of particles into 'fractions', according to how readily they sediment (Box 1). For example, at low centrifugation speeds, large cells can be separated from small cells. In the 1940s and '50s, advances in the technology of centrifugation allowed samples to be spun at very high speeds indeed, a refinement known as ultracentrifugation. For example, a centrifugation speed of 30 000 revolutions per minute (rpm) can produce a centrifugal force of 100 000 times Earth's gravitational force (100 000 × g). Ultracentrifugation using the appropriate types of sample preparation and solutions allows the separation of very small particles such as organelles and even macromolecules.
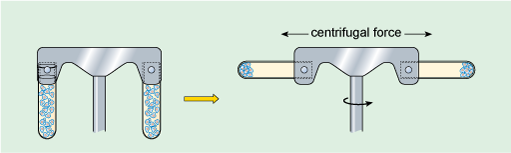
The left part of the diagram shows two vertical tubes on opposite sides of the rotor, each containing the same volume of suspension. On the right, the tubes are shown spinning and are now horizontal. The centrifugal force has caused the suspended material to form a pellet at the bottom of each tube.
Box 1 Cell fractionation
When a suspension of cells is placed in an appropriate physiological solution (an artificially prepared solution, which has pH and salt concentrations that mimic the internal environment of the cell) and disrupted, for example by detergents or ultrasound, or by homogenisation, many of the organelles such as nuclei, mitochondria and Golgi apparatus (Figure 1a and b) remain intact. The homogenisation apparatus consists of a tube with a tightly fitted pestle that is rotated to shear the cells (Figure 5a). The cell membrane and the endoplasmic reticulum are disrupted, but their membranes can re-seal to form vesicle-like structures called microsomes. If conditions are right, the original chemical properties of the organelles and membranes are retained during the preparation.
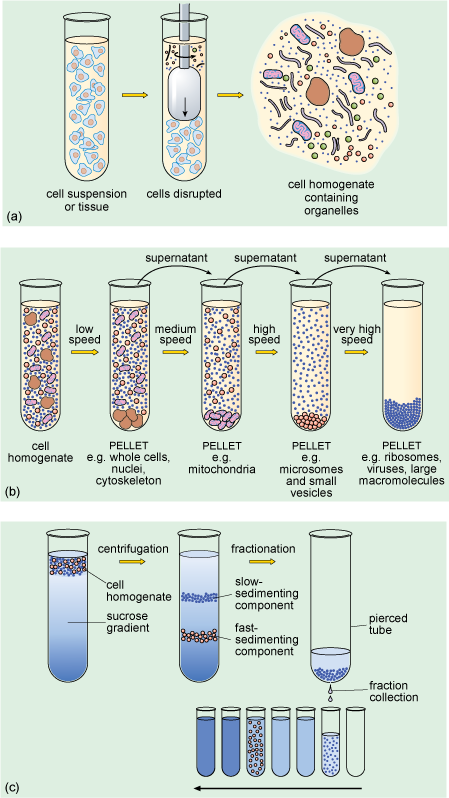
Part (a) shows that the disruption of the cells produces a homogenate comprising intact organelles and resealed membranes called microsomes. In part (b), low speed centrifugation of a cell homogenate produces a pellet of larger components such as whole cells, nuclei and cytoskeleton; medium speed centrifugation produces a pellet of smaller components such as mitochondria; high speed centrifugation produces a pellet of yet smaller components such as microsomes and small vesicles; and very high speed centrifugation produces a pellet of even smaller components such as ribosomes, viruses and large macromolecules. In part (c), centrifugation of a cell homogenate in a sucrose density gradient produces a fast-sedimenting, high density band of particles low down the tube and a slow-sedimenting, low density band of particles further up the tube. The tube is pierced at the bottom and successive fractions of decreasing density are collected.
The subcellular components can then be recovered by centrifugation. The cell homogenate is dispensed into centrifuge tubes which are placed into a rotating holder (known as a rotor) that fits into the centrifuge. As the rotor turns, particles suspended in the homogenate migrate towards the bottom of the tube. Particles collected at the bottom of the tube form a 'pellet'. Since larger components travel fastest, they can be separated from smaller particles by differential sedimentation (Figure 5b), in which centrifugation at progressively higher speeds is used to separate particles in the homogenate according to their size and density. After an initial centrifugation, the pellet, containing the largest components, is separated from the remaining suspension (known as the supernatant) which contains the smaller components. The pellet can be re-suspended and its components studied, while the supernatant can be transferred to another centrifuge tube and spun at a higher speed to recover smaller components into another pellet. A series of such steps can be performed to separate the different components of a cell homogenate.
A problem with differential sedimentation is that separation is not complete; pellets actually contain something of a mixture of cellular components. A more sensitive and widely used method is density gradient centrifugation in which the cell lysate is laid on top of a density gradient. This is prepared in a centrifuge tube by layering sucrose solutions of varying densities, the solutions becoming denser towards the bottom of the tube. Movement through the density gradient optimises the separation of particles of different density and size into different levels or bands in the gradient, so that they can be removed as separate 'fractions' (Figure 5c), either by using a fine pipette or by piercing the bottom of the tube and collecting the fractions as the liquid drips out.
Once separation is achieved, the biochemical properties of the different fractions can be analysed. Fractionation methods have been of great importance in cell biology because they enabled the functions of the different cell organelles to be elucidated. Indeed this method provided the first evidence for the existence of some organelles, such as lysosomes and peroxisomes (small membrane-bound organelles that contain different types of degradative enzymes, described later in this course).
Summary of Sections 1 and 2
- The internal structure of cells is studied by transmission electron microscopy, which allows cell organelles to be visualised.
- Immunoelectron microscopy allows the localisation of molecules to specific parts of cells.
- Cell fractionation techniques, including ultracentrifugation, allow cell organelles to be separated for subsequent biochemical analysis.
3 The organisation of prokaryotic cells
You have learnt that the general appearance of prokaryotic cells is relatively simple; but what about their internal organisation?
The interior of prokaryotic cells appears rather homogeneous when viewed with a light microscope because they do not contain a nucleus or membrane-bound organelles. When viewed under the electron microscope however, areas of different electron density can be seen. The prokaryotic genome typically consists of a circular DNA molecule located in the cytoplasm. The area in which the DNA is located (the nucleoid) appears relatively pale, whereas the ribosomes, where protein is synthesised, are darker, and give the cytoplasm a granular appearance, as can be seen in Figure 3. You will learn more about ribosomes in Section 4.4.
Some bacteria have visible cytoplasmic 'inclusions' that are not membrane-bound. Examples of some different inclusions are: gas vesicles (which are surrounded by a protein shell and have a role in the buoyancy of some aquatic bacteria), endospores (dormant structures formed within some types of bacteria when growth conditions are unfavourable, which can later germinate and give rise to new individual bacteria) and glycogen granules (glycogen is a polymer of glucose, which some bacteria store as an energy reserve).
A distinctive feature of bacteria, when viewed at either high or low magnification, is their surface. At high magnification, it can be seen that there are several distinct components to the structure that surrounds the bacterium. Like all other cells, bacterial cells are surrounded by a cell membrane, which is the innermost of these components. Most bacteria also have a cell wall that surrounds the cell membrane. This provides support, gives the cell its shape and prevents the cell from expanding, or even bursting by taking up too much water. The cell walls of Bacteria (but not of Archaea) are composed of peptidoglycan, which is a large polymer consisting of a complex arrangement of sugars linked by amino acids (including three amino acids that are not found naturally in proteins). The cell wall is different in Gram-positive and Gram-negative bacteria (Figure 6).
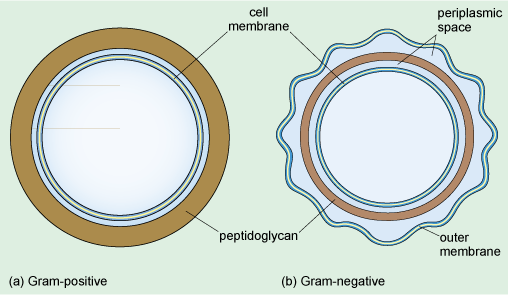
In both Gram-positive and Gram-negative bacteria the cytoplasm is enclosed by a cell membrane. Outside of this in the Gram-positive bacterium is a thick peptidoglycan outer layer. The Gram-negative bacterium has a much thinner peptidoglycan layer, either side of which is the periplasmic space. The outermost layer of the Gram-negative bacterium is the outer membrane.
From Figure 6, what is the difference between the cell walls of Gram-positive and Gram-negative bacteria?
The peptidoglycan layer is much thicker in Gram-positive bacteria.
The cell wall of Gram-negative bacteria consists of a much thinner layer of peptidoglycan forming a loose network within the periplasmic space separating the inner and outer membranes. The periplasmic space is filled with a gel and contains various enzymes and other proteins. It is the dense peptidoglycan layer that traps the purple Gram stain inside Gram-positive bacteria. In contrast, the thin peptidoglycan layer of Gram-negative bacteria is not sufficient to retain the stain, and in the final step of the Gram staining procedure, treatment with an organic solvent dissolves the cell membranes and decolourises the cell. Gram-negative bacteria have a second membrane known as the outer membrane (Figure 6) which is outside the cell wall.
Finally, some bacteria secrete polymers, usually polysaccharides and sometimes proteins, which form a slimy, outer capsule that protects the cell from desiccation and from phagocytosis.
Some bacteria have internal specialisations: for example, in the photosynthetic cyanobacteria, the cell membrane folds into the cytoplasm to form stacks of membranes, which are the site where photosynthesis occurs. In other prokaryotes, the cell membrane folds inwards in a more irregular arrangement. Some bacterial species also have structures extending outwards from the cell membrane from the cell membrane, known as flagella and pili.
Summary of Section 3
- Prokaryotes do not have a nucleus or other internal membrane-bound organelles and their cytoplasm appears relatively homogeneous under the light microscope; their DNA appears pale, because the rest of the cell is densely packed with ribosomes, which are far more electron-dense and so appear dark.
- Most bacteria have a cell wall that lies outside the cell membrane. The cell wall of Gram-positive bacteria is composed of a thick layer of peptidoglycan, while Gram-negative bacteria have a thinner layer of peptidoglycan within the periplasmic space between the inner and outer cell membranes.
- Some bacteria possess membrane specialisations (relatively common examples are flagella and pili) and a few exhibit complex folding of the cell membrane.
4 The organisation of eukaryotic cells
In eukaryotic cells many activities are compartmentalised within the organelles. The different organelles serve different functions, although in fact each type of organelle (e.g. the nucleus, or the endoplasmic reticulum, described later in this section) may play a role in several different activities. Some activities, for example the production and processing of proteins, involve different parts of the cell, or several organelles, as you will see.
Suggest some key cellular functions or processes.
You may have thought of the synthesis of different macromolecules (DNA, proteins, lipids and carbohydrates), secretion, movement, cell division, protection, and there are many more!
Below, a 'tour' through the eukaryotic cell describes the different subcellular components and their functions, with an emphasis on a typical higher (mammalian) animal cell, but also describing some of the cell components in other eukaryotic organisms. It should become clear, as you work through this section, that eukaryotic cells and their components are highly dynamic.
4.1 Cell surfaces
The cell surface is functionally very important, because it is the cell's interface with its environment.
Suggest two functions of the cell surface.
Protection, and absorption of nutrients. You may also have thought of: secretion of signalling molecules or enzymes, disposal of cellular wastes, gas exchange, or cell-cell recognition.
One major difference between different eukaryotes is the presence or absence of a cell wall.
In which eukaryotic kingdom do all cells lack a rigid cell wall?
The cells of organisms in the animal kingdom lack a rigid cell wall (Figure 1a).
Plant cells (Figure 1b) and fungal cells have a cell wall, which provides support and determines cell shape, while protists are diverse in this respect; some, such as amoebae, do not have a rigid cell wall, but many do. All cells, however, have a cell membrane, which acts as a barrier and an interface with the external environment.
The cell membrane is composed predominantly of phospholipids, arranged in a bilayer. A schematic diagram illustrating the main features of a typical animal cell membrane is shown in Figure 7. Within the bilayer are embedded a variety of proteins and glycoproteins (proteins that have sugars attached). These proteins, many of which span the membrane, play crucial roles in the interactions of the cell with its environment. Some membrane proteins act as transporters or channels that allow selective movement of ions, nutrients or other molecules into the cell. Others, often glycoproteins, act as receptors, which respond to specific molecular changes in the extracellular environment and transduce information about the environment to the inside of the cell, thereby allowing appropriate responses to be initiated.
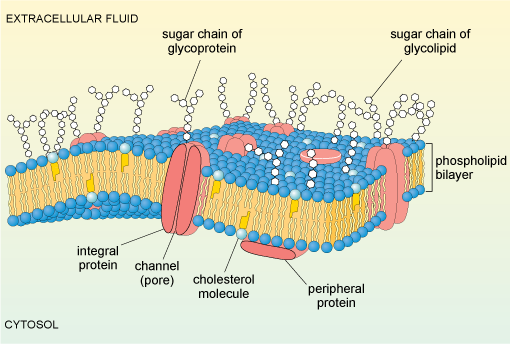
The diagram shows the double layer of phospholipid molecules: the polar head of each phospholipid molecule is shown as a blue sphere and the two hydrophobic or non-polar tails attached to the head are shown as zig-zag yellow ribbons. The tails in each layer are oriented in parallel and are facing, but not touching, the tails of the opposing layer in the bilayer structure. Interspersed between the phospholipid molecules are so-called integral protein molecules (shown here in red) which span the width of the membrane and provide aqueous channels or pores in the membrane. Other protein molecules, the peripheral membrane proteins, are attached to the cytosolic surface of the membrane. Some of the proteins and lipids at the extracellular surface of the membrane have short, sometimes branched sugar chains attached; these are called glycoproteins and glycolipids, respectively.
Yet other glycoproteins act as recognition molecules, and can promote adhesion between adjacent cells in a tissue. The cell membrane is also linked to proteins on its cytoplasmic surface (intracellular proteins). These include components of the cytoskeleton, which have a structural role, maintaining the shape of the cell.
4.2 The cytoskeleton
The cytosol of eukaryotic cells contains a system of specialised protein assemblies that together form what is known as the cytoskeleton. Another term often used is cellular 'scaffolding' which is perhaps rather misleading, since the protein assemblies that make up the cytoskeleton are not fixed, but are highly dynamic and play an essential role in the transport of organelles and some molecules within the cell. Cytoskeletal proteins also have other roles; their actions produce all kinds of cell movements, including the movement of motile cells and intracellular movements, such as those of chromosomes during cell division. In animal cells, the cytoskeleton also provides mechanical strength and support, and helps to maintain cell shape. In plant cells, the cell wall fulfils this role.
It used to be thought that prokaryotic cells did not have a cytoskeleton, but recent research has revealed that they have filaments assembled from proteins similar to those found in eukaryotic cells, e.g. actin. The prokaryotic cytoskeleton has roles in cell division and maintaining cell shape and polarity.
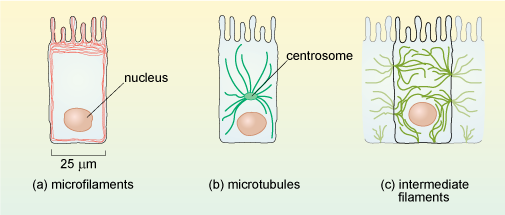
Cytoskeletal proteins form long, filament-like assemblies. There are three types of filaments, formed from different proteins: microfilaments (also known as actin filaments), microtubules and intermediate filaments (Figure 8).
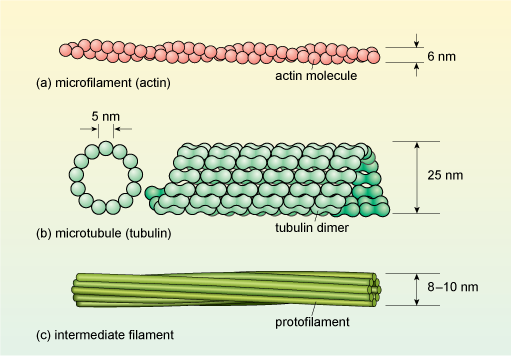
In (a) the microfilament comprises two chains of actin monomers in a loose spiral structure the diameter of which is about 6 nanometres. Part (b) includes a schematic cross-section through a microtubule which is a ring of 13 tubulin monomers each of diameter about 5 nanometres; there is also a side-on view of part of a microtubule, showing the dumb-bells-shaped tubulin monomers arranged in a spiral structure to form the tubule. The external diameter of the tubule is about 25 nanometres. The intermediate filament shown in part (c) comprises about eight narrow protofilaments, which do not have a subunit structure. The protofilaments are wound around each other in a rope-like structure 8-10 nanometres in diameter.
Microfilaments
Each type of cytoskeletal protein has a distinct distribution in cells, as illustrated in Figure 9 for intestinal epithelial cells. All three types of filaments are associated with other types of protein in the cell.
Microfilaments are present in all eukaryotic cells and are the thinnest of the filament types, having a diameter of about 6 nm (Figure 8a) and a thread-like appearance under EM. They are composed of molecules of the protein actin organised into a long helical chain, and so are also sometimes called actin filaments. Most microfilaments do not occur singly but are linked together in networks or bundles.
Networks of microfilaments are particularly prominent around the edges of cells, in a region just below the cell membrane, sometimes called the cell cortex (Figures 9a and 10). Bundles of microfilaments are found in microvilli of absorptive epithelial cells (e.g. in the intestine), and in the leading edge of moving cells, where the ability of the actin filaments to rapidly disassemble and reassemble plays a key role in cell motility.
Microtubules
Microtubules are, as their name implies, tubular structures composed of two forms (called α and β) of a protein known as tubulin. They are present in all eukaryotic cells and play a crucial role in maintaining cell shape and also in intracellular movement: for example, the movement of cell organelles from one part of the cell to another, and the reorganisation of chromosomes into the daughter cells during cell division. Microtubules are effectively hollow tubes typically consisting of 13 parallel filaments of tubulin assemblies, and they measure about 25 nm in external diameter (Figure 8b).
In animal cells, microtubules radiate from a microtubule organising centre (MTOC) known as the centrosome which is located near the nucleus (Figure 1a and Figure 9). Typical plant cells do not have an equivalent of the centrosome. Instead, less well-defined microtubule organising centres (MTOCs) appear at different times and places to organise microtubule networks in different types of plant cell. Microtubules are very unstable, and are constantly being disassembled and reassembled. They begin to assemble at the centrosome, and the growing tubules extend out radially, towards the edges of the cell (Figure 10). Some microtubules disassemble before they reach the cell cortex, but they will become stabilised if they attach to an organelle or bind to a protein known as a capping protein.
What might happen to the shape of a cell if the capping proteins become confined to one part of the cell cortex?
Microtubules would only become stabilised at that region of the cell periphery, so the cell could become polarised; that is, it could acquire an asymmetrical shape.
What types of animal cell are inherently asymmetrical in shape?
Neurons and some kinds of epithelial cells are examples of asymmetrical cells .
The selective stabilisation of microtubules, determined by the location of capping proteins, is therefore an important factor in determining cell shape. The arrangement of microtubules also plays an important role in the distribution of organelles within the cell.
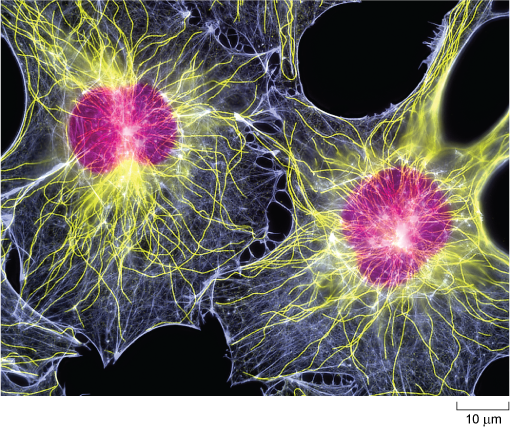
In this micrograph, the near-spherical, purple-fluorescing nuclei are near the centre of the cells, both of which have an irregular webbed shape. The cytoplasm is filled with a mesh of very fine blue-fluorescing microfilaments and also thicker yellow-fluorescing microtubules, which appear to radiate out from the nucleus like strands of cooked spaghetti.
Intermediate filaments
Intermediate filaments are so called because they are intermediate in diameter between microfilaments and microtubules, measuring about 8-10 nm. Whereas microfilaments and microtubules are both assembled from a single type of protein (actin and tubulin, respectively), there are several types of intermediate filament protein. You are probably most familiar with the type called keratins, which are abundant in the cells at the surface of mammalian skin. An intermediate filament contains about eight 'protofilaments' wound around each other in a rope-like structure (Figure 8c) which confers great strength. Unlike actin filaments and microtubules, the intermediate filaments are not involved in cell movements; their main role is to provide mechanical strength to cells and tissues.
4.3 The nucleus
The nucleus is the largest, and so usually the most easily identified cell organelle. In a typical animal cell, the nucleus is roughly spherical in shape (Figure 11) and is often, but not always, situated near the centre of the cell. The nucleus contains the majority of the genetic information of the cell. Two of the major activities that take place in the nucleus are: DNA replication (the synthesis of new DNA in preparation for cell division) and transcription (the production of RNA copies of parts of the DNA sequence). The production of a messenger RNA (mRNA) is the first step in the synthesis of proteins. Another important process that takes place in the nucleus is the assembly of ribosomes, the complex structures in the cytoplasm where mRNA is translated into protein.
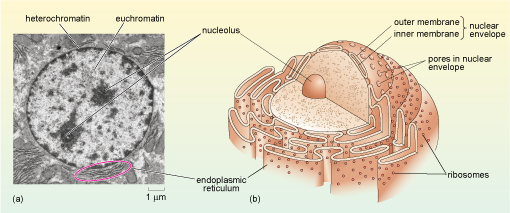
The diagram in part (b) shows the nucleus surrounded by the nuclear envelope consisting of an inner and an outer membrane and studded with nuclear pores. Immediately outside the nuclear envelope is the rough endoplasmic reticulum which has ribosomes on its outer (i.e. cytosol-facing) surface.
In order for these three major activities to take place, much coordination and many other 'subsidiary' activities are necessary. For example, replication and transcription do not take place at the same time. The enzymes needed to synthesise DNA or transcribe it into RNA must be present at the right places and times, and, since protein synthesis occurs in the cytoplasm, there must be a means of exporting mRNA (and ribosomes) out of the nucleus, and specific proteins into it. How does the organisation of the nucleus and its component molecules allow all of these activities to take place?
The arrangement of DNA in the nucleus
You will recall that in eukaryotes, DNA is organised into chromosomes, each of which contains a single, very long double-stranded DNA molecule. These molecules are much longer than the diameter of the nucleus, and are packaged into the nucleus by winding around proteins known as histones, which help to coil the DNA, as shown in Figure 12. This complex of DNA and proteins is known as chromatin. The binding of DNA to histones is very ordered and the degree of packing varies. The DNA is most tightly condensed in mitotic chromosomes (as shown in Figure 12) just before cell division, when the highly condensed chromosomes can be easily seen under the light microscope. The rest of the time, individual chromosomes are not visible by light microscopy because they are far less condensed, allowing easy access to the enzymes that carry out DNA replication and transcription. Constant adjustment of DNA packaging and the spacing between nucleosomes is one way in which gene expression is controlled.
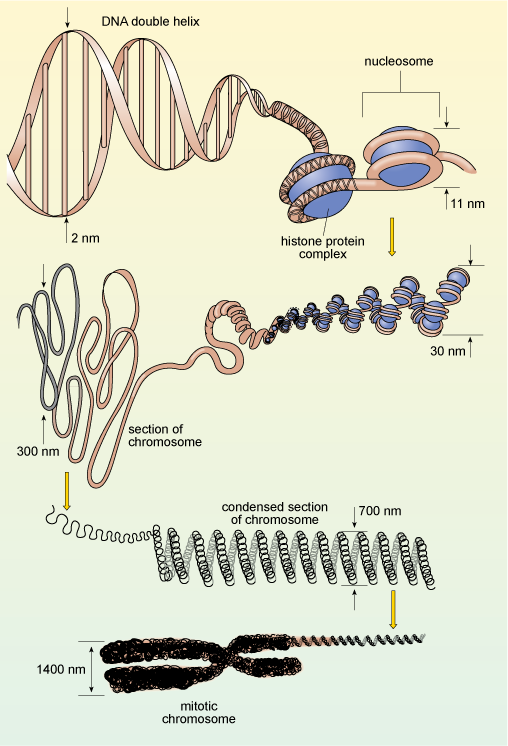
The diagram shows the dimensions of the chromosome components from the bare, unfolded DNA double helix of diameter 2 nanometres to the highly compacted mitotic chromosome. The double helix is wound around a histone-protein complex, forming a succession of nucleosomes - an extended chromatin strand of width about 11 nanometres, The strand is wound into a spiral structure of width 30 nanometres, which is looped and condensed into a structure 300 nanometres wide, characteristic of cells during interphase. Further spiralling and condensation occurs in cells preparing to divide, when each member of a pair of chromatids has a diameter of about 700 nanometres.
If you look at Figure 11a, you see that some areas of chromatin (particularly around the edges of the nucleus) have a dark, electron-dense appearance, while other areas of the nucleus appear paler. The chromatin in the dark areas is known as heterochromatin and is more highly condensed than the DNA in the paler areas which is called euchromatin. In general, heterochromatin contains far fewer genes and is far less 'transcriptionally active' than euchromatin.
The nucleolus
Another example of the way that the organisation of chromosomes within the nucleus relates to function is in the nucleolus (plural, nucleoli), where ribosomes are assembled. When viewed by electron microscopy, nucleoli appear as large rounded patches of an electron-dense material with a granular appearance (Figure 11a). Ribosomes are composed of protein and RNA molecules (Section 4.4) and the genes needed for the production of the ribosomal RNA components are located in clusters on several of the chromosomes. The relevant sections of these chromosomes loop into the nucleolus (Figure 13), where the ribosomal RNA molecules transcribed from this DNA are packaged together with ribosomal proteins imported from the cytosol. The assembled ribosome subunits are responsible for the electron-dense appearance of the nucleoli in TEM.
The size of the nucleolus reflects the activity of the cell.
Given the function of the nucleolus, what might its size indicate about the activity of a cell?
The presence of a large nucleolus (or multiple nucleoli as in Figure 11a) suggests that the cell is synthesising a large amount of protein, as the nucleolus is the site of ribosome assembly, and ribosomes are required for protein synthesis in the cytosol. A small nucleolus suggests that a cell is not synthesising much protein.
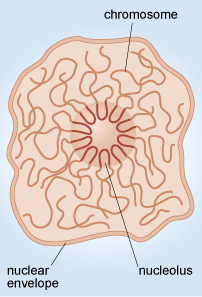
This diagram shows the nucleolus at the centre of the nucleus. A short region of each of the chromosomes forms a loop inside the nucleolus while the remainder of each chromosome extends outwards, into the main body of the nucleus.
Nuclear structure and the transport of molecules
The structure of the nucleus is maintained by a family of intermediate filament proteins known as lamins, which form a network of filaments on the inner surface of the nuclear membrane. This network is called the nuclear lamina and is linked both to the nuclear membrane and to the chromatin; the nuclear lamina is thought to help to organise the chromatin.
When viewed at high magnification, it can be seen that the membrane that surrounds the nucleus is actually a double membrane, or nuclear envelope, as it is sometimes called. Within the envelope are gaps called nuclear pores (Figure 11b and Figure 14) which create channels of about 9 nm diameter. Small water-soluble molecules diffuse freely through the nuclear pores, but the movement of proteins into and out of the nuclear pore is regulated by a complex arrangement of proteins known as the nuclear pore complex. The nuclear pore complex allows RNA molecules and ribosomes to pass out of the nucleus, and allows selected proteins to enter the nucleus, but prevents passage of most other proteins.
What type of proteins would you predict to be transported into the nucleus from the cytoplasm?
Histones, which are needed for the packing of DNA; the enzymes and other proteins needed for replication and transcription of DNA; the proteins needed for ribosome assembly; and the proteins that make up the nuclear lamina.

This electron micrograph shows a small section of the nuclear envelope with numerous circular pores about 10 nanometres in diameter. The spacing between the pores is between 10 and 15 nanometres.
The nucleus, then, is crucial as the site of DNA replication, transcription of DNA into RNA and ribosome assembly. You have seen that the ribosome subunits and mRNA molecules leave the nucleus through the nuclear pores. The next section describes what happens to them in the cytosol.
4.4 Ribosomes: the sites of protein synthesis
The mRNA molecules exported from the nucleus are translated into protein in the cytoplasm by the ribosomes (which are RNA-protein complexes, not organelles). Evidence suggests that some mRNA molecules leaving the nucleus are directed to specific sites in the cell before they are translated. Ribosomal structure is similar in prokaryotic and eukaryotic cells. Ribosomes are organised into a large subunit and a small subunit, as shown in Figure 15 (prokaryotic ribosomes contain over 50 different types of protein and those of eukaryotic cells contain more than 80 different types of protein). The large subunit contains two different ribosomal RNAs in prokaryotes and three in eukaryotes, while the small subunits contain one ribosomal RNA'.
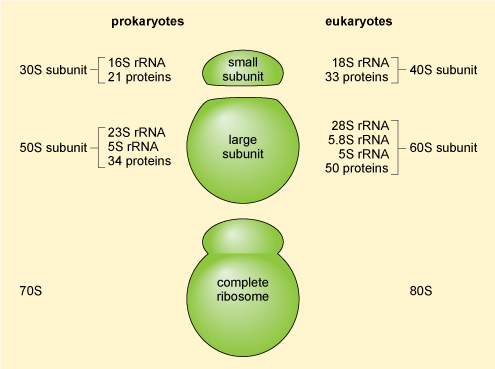
This diagram summarises the structural organisation of prokaryotic and eukaryotic ribosomes. Prokaryotic ribosomes are 70S particles comprising a small, 30S subunit and a large, 50S subunit. The 30S subunit is made up of 16S rRNA and 21 proteins; and the 50S subunit is made up of 23S rRNA, 5S rRNA and 34 proteins. Eukaryotic ribosomes are 80S particles comprising a small, 40S subunit and a large, 60S subunit. The 40S subunit is made up of 18S rRNA and 33 proteins; and the 50S subunit is made up of 28S rRNA, 5.8S rRNA, 5S rRNA and 50 proteins.
Ribosomes occur both 'freely' in the cytosol, i.e. not attached to any organelle (Figure 16), or attached to the membrane of the rough endoplasmic reticulum in eukaryotic cells (Figure 1a and b). Cytosolic proteins and proteins that are destined for the nucleus, mitochondria, chloroplasts and peroxisomes (you will learn about these other organelles later in this course) are synthesised by the free ribosomes in the cytosol. Cytosolic proteins, such as cytoskeleton components, are simply released into the cytosol once their synthesis is complete, but many other proteins are delivered to specific locations by a complex system of protein targeting or protein sorting. A signal sequence, a characteristic short sequence of particular amino acids in the protein itself, acts rather like a postal address that targets the protein for import into the correct organelle. Nuclear proteins, for example, contain a nuclear localisation sequence and are recognised by proteins that transport them to the nuclear membrane and through the nuclear pore. Proteins destined for mitochondria, chloroplasts or peroxisomes also have a characteristic signal sequence directing them to the appropriate organelle. This 'tagging' of polypeptides by signal sequences comprising specific amino acids is an important general mechanism used by all cells to recognise and deliver proteins to specific sites in the cell. Prokaryotic cells lack organelles, but they use a similar mechanism to target proteins to the cell membranes, cell wall, or to various types of inclusions in the cytosol.
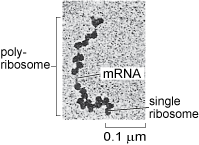
As you will learn in the next section, lysosomal proteins (Section 4.9), and proteins that are destined for export from the cell, or to reside in the cell membrane, are translated by ribosomes that are attached to the endoplasmic reticulum.
4.5 The endoplasmic reticulum
The nuclear membrane is continuous with an extensive membrane system known as the endoplasmic reticulum (ER, see Figure 1 and Figure 11), which extends through much of the cytoplasm and is the site of many activities, including the synthesis of lipids in the smooth endoplasmic reticulum and the synthesis of lysosomal proteins, cell membrane proteins and secreted proteins in the rough endoplasmic reticulum. Importantly, the interior of the ER is separate from the cytosol. The membranes of the ER form tubes and sacs, and depending on the plane in which a section cuts through the cell, the ER may therefore have the appearance of circular or elongated tubes, or parallel systems of membrane, or any shape in between, as illustrated in Figure 17.
Some parts of the endoplasmic reticulum appear smooth when viewed by EM and are hence named smooth endoplasmic reticulum (SER). Smooth endoplasmic reticulum is involved in the production of phospholipids and steroids and also in the detoxification of substances such as drugs or ingested chemicals. Ingestion of large amounts of a toxic substance, such as a pesticide, results in an increase in the amount of SER in cells. Normally, however, most of the endoplasmic reticulum appears granular, being dotted with many ribosomes (Figure 17). This part of the endoplasmic reticulum is given the name rough endoplasmic reticulum (RER), and is the site where lysosomal proteins (mainly digestive enzymes), membrane proteins, and proteins that are destined for export from the cell are synthesised and processed.
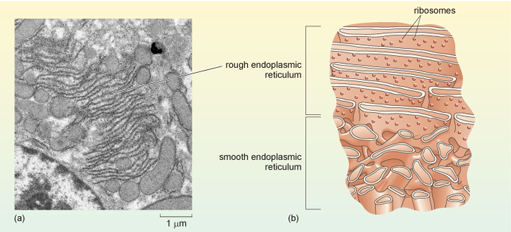
The electron micrograph in (a) shows part if the nucleus (bottom left) and multiple layers of rough endoplasmic reticulum membranes. The diagram in part (b) is a 3D cross-section showing both the ribosome-studded rough endoplasmic reticulum and adjacent smooth endoplasmic reticulum.
What determines whether a particular mRNA molecule is translated by ribosomes that are free in the cytosol or by ribosomes that are attached to the RER? The answer lies again in the amino acid sequence of the protein being synthesised. In fact, translation of all eukaryotic mRNAs starts on 'free' ribosomes in the cytosol (Figure 18a), but if an mRNA encodes a protein that is destined for lysosomes, or to be secreted from the cell, or to be embedded in a cell membrane, the ribosome is rapidly redirected to the endoplasmic reticulum by a process known as cotranslational localisation. As soon as the beginning of the translated polypeptide (which is known as the N-terminus, or amino (NH2) terminus) starts to emerge from the free ribosome, a signal sequence for localisation to the RER, close to the N-terminus, is recognised by a protein complex called the signal recognition particle (SRP). Translation pauses, and the whole complex, including the ribosome, mRNA and partially translated polypeptide, is transported to the membrane of the RER. There, translation recommences and the growing polypeptide chain is simultaneously translocated across the membrane of the RER and starts to enter the RER lumen, the space inside the RER (Figure 18c).
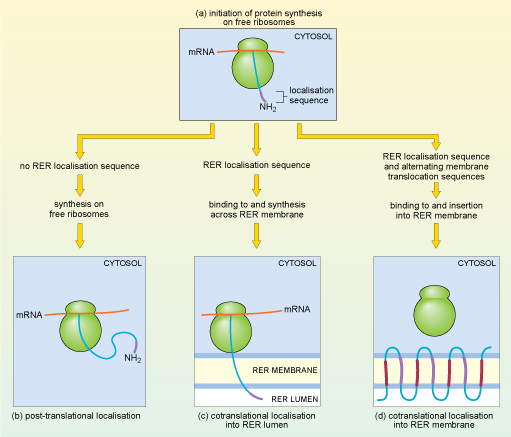
One of two things may then happen: for some proteins, the translated polypeptide continues to enter the RER lumen, until finally the completed polypeptide is released inside the RER. These proteins usually pass on to the Golgi apparatus (see Section 4.7) and are eventually secreted from the cells or packaged into lysosomes. Alternatively, some proteins remain partly embedded within the RER membrane.
Thinking of how polypeptides are 'directed' to the RER, suggest a molecular mechanism that would determine whether a particular protein remains in the RER membrane, or enters the RER lumen.
A protein that is destined to remain in the RER membrane could have a special 'stop' sequence of amino acids, which remains in the membrane and thus prevents the protein from completely entering the lumen.
That is indeed what happens; a stop-transfer signal halts translocation and the protein remains embedded in the ER membrane. These proteins are either destined to be delivered to other membranes, usually the cell membrane, or remain in the ER, where they play a role in ER function. Some proteins, many of them receptor proteins, are 'multipass' transmembrane proteins. One example, rhodopsin, is a light-sensitive receptor protein involved in light perception in the retina of the mammalian eye. It has seven membrane-spanning sections or domains. A signal sequence at the N-terminus of the rhodopsin protein acts as a start-transfer signal which directs translocation of the growing polypeptide chain through the ER membrane until a stop-transfer signal is reached (Figure 18d). A second start-transfer signal sequence further along the growing polypeptide chain threads another section of the protein across the membrane until it reaches a second stop-transfer signal. Alternating start- and stop-transfer signals allow the rhodopsin protein to thread back and forth through the membrane seven times.
The proteins translated in the RER are then modified in different ways. One important modification that begins in the RER is protein glycosylation; that is, the addition of sugar residues; glycosylated proteins are known as glycoproteins (Figure 7). Some sugar residues can also serve as part of the 'address label', and are involved in protein targeting later on; others have different roles. The initial glycosylation of proteins occurs in the RER, through the action of enzymes that are embedded within the RER membrane. The modified proteins that are destined for export or to be delivered to lysosomes or the cell membrane, then pass from the RER to the Golgi apparatus (see Section 4.7), for further processing.
4.6 Vesicles
The proteins and glycoproteins leaving the RER are transported to the Golgi apparatus in small spherical membrane-bounded compartments, known as transport vesicles, which pinch off from the RER. The membrane of the transport vesicle fuses with the Golgi membrane, so proteins inside the vesicle are released into the Golgi lumen while proteins embedded in the vesicle membranes are incorporated into the Golgi membrane. After further processing, sorting and packaging in the Golgi apparatus, the modified proteins are again packaged into vesicles (Figure 19b) and delivered to their final destination in a similar manner. Secretory cells usually have large numbers of secretory vesicles which fuse with the cell membrane to release their contents outside the cell. There are many different types of vesicle, even in cells that are not specialised for secretion. Vesicles are also used to excrete waste materials and to transport many different molecules within cells, for example proteins and glycoproteins. Newly synthesised lipids from the SER are incorporated in vesicle membranes and are therefore also delivered to the cell membrane in these small structures.
How do vesicles move within cells, and what mechanism ensures their correct delivery? The surfaces of vesicles have different coatings and surface signals that identify their contents and ensure that they fuse with the correct target membrane. Vesicles can move short distances within the cytosol by diffusion, but movement to more distant sites in the cell is mediated by the action of special 'motor' proteins that are able to 'walk' vesicles along the cytoskeleton network.
4.7 The Golgi apparatus
The main functions of the Golgi apparatus are: to sort the proteins and lipids arriving from the ER according to any 'labels' they carry; to complete the glycosylation (and some other modifications) of proteins; and finally, to package the proteins and lipids into vesicles according to one of three main destinations: (1) the lysosomes, (2) the cell (plasma) membrane, or (3) secretion from the cell.
When viewed in the electron microscope, the Golgi apparatus appears as stacks of smooth, flat membranous sacs, known as cisternae, which are slightly curved and surrounded by vesicles of various sizes (Figure 19). The Golgi often has a relatively inconspicuous appearance compared to the RER and its size varies in different cell types.
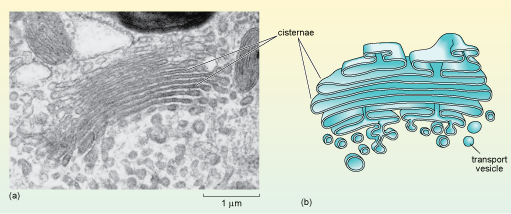
The diagram in part (b) is a 3D cross-section showing the cisternae with their slightly bulging ends, and numerous transport vesicles budding off from the lowermost cisternae.
Why might the size of the Golgi be different in different types of cells?
Different cells secrete different amounts of proteins and lipids. Some cells, such as enzyme-secreting cells, for example, secrete large quantities of these proteins and so are specialised for their production and packaging for export. Other types, such as smooth muscle cells, export very little protein.
Would you expect a cell that does not secrete any lipids or proteins to have a Golgi apparatus?
Yes, all cells need at least small quantities of lipid and protein to be delivered to their cell membrane. So, all eukaryotic cells have a Golgi apparatus.
4.8 Protein secretion from the cell
Intestinal epithelial cells are a good example of the diversity of secretory function. The intestinal epithelium contains many cells that are specialised for absorption (and have a small Golgi apparatus and only a few secretory vesicles), but it also includes several different types of cell that are specialised for secretion. Some of these cells secrete digestive enzymes from their apical surface which is exposed to the intestinal lumen (where ingested food is located); different enzymes are secreted in different parts of the gastrointestinal tract. Cells of another type secrete heavily glycosylated proteins known as proteoglycans into the lumen of the intestine. These mix with water to form mucus which acts as a lubricant, aiding the passage of material along the gut. Yet other epithelial cells secrete local hormones, from the other surfaces of the cell (the basolateral sides. i.e. base (basal) and side (lateral)) that are in contact with other cells. These hormones are secreted either into blood vessels or close to the surface of nearby cells.
Each type of intestinal epithelial cell is therefore polarised; it is not uniform, either in shape or function, as shown schematically in Figure 20.
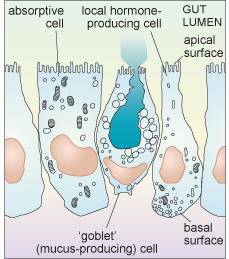
Another factor that is important to think about when considering the secretion of molecules from cells is the difference in demand for different types of exported molecules. For example, some substances are constantly delivered to the cell membrane, and are continuously released in small quantities all the time. This sort of release is known as constitutive secretion. An example is the secretion of mucus from intestinal epithelial cells. Other molecules, however, are only released at certain times, in response to some kind of signal. This type of release is known as regulated secretion. An example of regulated secretion is that of gastrointestinal hormones and digestive enzymes, which are released in response to ingestion of food.
Within each of these three types of intestinal epithelial cell, a complex sorting and delivery system exists. All the membrane-associated and secretory proteins are synthesised in the RER, processed and packaged in the Golgi, and then they continue by transport in vesicles along the cytoskeleton to the cell membrane. What about the mechanism by which molecules are actually released from cells? This involves a process known as exocytosis. During exocytosis the vesicle membrane fuses with the cell membrane, releasing the vesicle contents outside the cell.
What is the general term for the converse process, in which extracellular materials are ingested by engulfment by extensions of the cell membrane which form into vesicular structures within the cytosol?
The term is endocytosis.
You have already learnt that organisms such as amoebae feed by endocytosis (sometimes referred to as phagocytosis when it involves larger particles such as bacteria). Cells of the immune system, known as phagocytes, also ingest bacteria in this way. What is the fate of the materials that the cell ingests? They are broken down by yet another type of organelle, the lysosome.
4.9 Lysosomes and peroxisomes
Lysosomes are small spherical organelles, enclosed by a single membrane, which are common in animal cells but rare in plant cells. They measure about 0.5-1.0 µm across, and they contain digestive enzymes. Lysosomes fuse with membrane-bound endosomes (containing nutrients ingested by endocytosis), and the lysosomal enzymes digest large nutrient molecules. Cells can also degrade and recycle the components of their own organelles and structures when they are old or damaged, or if the cell is 'starving' in the absence of nutrients. This process, known as autophagy, usually involves formation of a membrane around the cell component and fusion of the resulting vesicle with lysosomes. Autophagy is thought to have an important role in many processes including cell growth, cell death and infection. Lysosomes contain many different kinds of digestive enzymes, and the inside of the lysosome is acidic, with a pH of around 5. The enzymes of the lysosome are specialised to perform their function only at this low pH; so, should they leak out into the cytosol, which has a pH of around 7.2, they do not do a great deal of damage. The inside of the lysosome is made acidic by the action of specialised transport proteins that lie in the lysosomal membrane and 'pump' hydrogen ions into its lumen. Other lysosomal membrane proteins transport the useful products of digestion out of the lysosome into the cytosol.
What useful materials would be produced by digestion in the lysosome?
The products would depend on the starting material, but could be amino acids, sugars and nucleotides.
Peroxisomes are also small enzyme-containing organelles bound by a single membrane, which are very similar in size to lysosomes, measuring between 0.2 and 1.0 µm in diameter. They are thought to be present in all eukaryotic cells. In mammals, they are particularly plentiful in liver cells and adipocytes but are much less abundant in other cells. Like lysosomes, peroxisomes also have a role in metabolism; they contain enzymes that break down fatty acids and amino acids, resulting in, among other things, the production of the toxic substance, hydrogen peroxide. Peroxisomes therefore also contain high levels of an enzyme known as catalase which breaks down the hydrogen peroxide into harmless products (water and oxygen). Lysosomes and peroxisomes are shown in Figure 21.
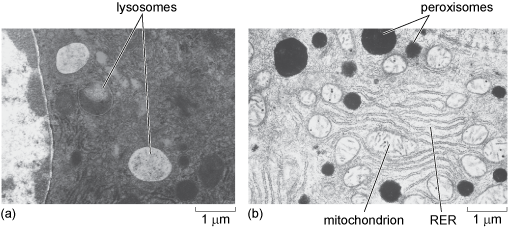
In the electron micrograph in (a), the lysosomes appear as pale (electron-lucent), almost circular structures of a range of sizes. In (b), the peroxisomes appear as very dark (electron-dense) circles of a large range of sizes. Also present in this image are a number of much less darkly staining mitochondria and some rough endoplasmic reticulum.
For many years, peroxisomes were thought to be identical to lysosomes in their properties. The difference between these organelles was discovered as the result of cell fractionation experiments (Box 1). Although very similar in size, their contents and therefore their densities are different, so under particular centrifugation conditions the two organelles sediment in different fractions. Different enzymes were found to be associated with the two fractions and it is now known that the two organelles are very distinct.
What is the difference between the site of synthesis of lysosomal and peroxisomal proteins?
Lysosomal proteins are synthesised at the RER (Section 4.5); peroxisomal proteins are synthesised by free ribosomes in the cytosol (Section 4.4).
How are the correct proteins delivered from the cytosol to the peroxisomes?
Peroxisomal proteins have a signal sequence to ensure their correct targeting.
Correct targeting of peroxisomal proteins to the peroxisomal membrane or interior occurs because of specific sequences in the proteins, as described previously.
Having studied the organelles and other cellular components involved in the synthesis and delivery of proteins in the cell, in the next section you will consider an organelle that you already know something about, the mitochondrion, which plays a fundamental role in the generation of ATP in eukaryotic cells.
4.10 Mitochondria
Mitochondria have a distinctive appearance when viewed by electron microscopy. They often appear as rounded or sausage-shaped structures (Figure 1a, b and Figure 22a, b), measuring about 0.5-1.0 µm in diameter and 2-8 µm in length; although their size and shape vary, and they are often much bigger in plants. For many years all mitochondria were assumed to have this fairly regular shape, but recent research, in which mitochondria have been observed in living cells, has shown that mitochondria are dynamic; they divide and fuse, and the shapes of individual mitochondria change. Mitochondria also move; their positions in the cell are not fixed. In many cells, mitochondria are long and extended and have a network-like structure (Figure 22c). So, rather like the ER, the appearance of mitochondria as round or elongated and flattened spheres in micrographs is often simply due to the plane in which they are sectioned.
Mitochondria have a double membrane; the inner membrane is thrown into many folds known as cristae, which are what gives the organelle its distinctive appearance. The two membranes effectively create two compartments in the mitochondrion. The intermembrane space lies between the two membranes; the much larger mitochondrial matrix lies within the inner membrane, as shown in Figure 22b.
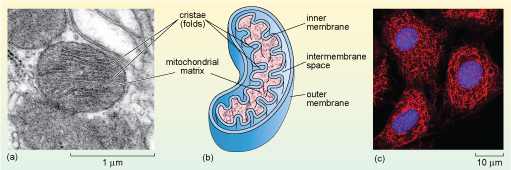
The schematic diagram in (b) shows the mitochondrion bounded by an outer membrane, inside which is a compartment called the intermembrane space, which separates the outer membrane from the highly folded inner membrane. The folds, or cristae, extend into the inner compartment of the organelle, called the mitochondrial matrix.
The mitochondrial matrix contains many enzymes, including some of those involved in the breakdown of glucose. It also contains mitochondrial DNA, and the molecules and cellular components needed to transcribe and translate mitochondrial genes including ribosomes. Mitochondrial genes encode most of the proteins of the inner mitochondrial membrane, which plays a fundamental role in the generation of ATP. The majority of mitochondrial proteins, however, are encoded by the cell's nuclear DNA and are synthesised in the cytosol and imported into the mitochondrion. Again, the correct delivery of mitochondrial proteins from the cytosol depends on specific signal or targeting sequences.
Mitochondria produce most of the ATP in eukaryotic cells, and are hence more abundant in cells that have high requirements for energy, such as muscle cells.
4.11 Chloroplasts
Chloroplasts, the sites of photosynthesis, are found in plant cells and photosynthetic algae. Like mitochondria, chloroplasts have their own ribosomes and their own DNA, which encodes the proteins of the internal membrane that use light energy to generate ATP in photosynthesis. Most other chloroplast proteins are, however, encoded by the nuclear DNA and must be transported into the chloroplasts. Chloroplasts are typically about 2-4 µm wide, and 5-10 µm in length and have three membrane systems.
In addition to an outer and inner membrane (which encloses a space filled by a fluid substance known as the stroma, described below), chloroplasts have a complex structure of internal membranes (here referred to as lamellae; singular, lamella). The lamellar structure consists of flattened sacs called thylakoids, which consist of a thylakoid membrane surrounding a thylakoid lumen (Figure 23a and b). The thylakoids are often stacked into grana (singular granum), which look rather like stacks of pancakes. The grana are linked by stroma lamellae, which join the stacks of grana together to form a single compartment. The thylakoid membranes and stroma lamellae have embedded within them the chlorophylls and other light-absorbing pigments required for photosynthesis.
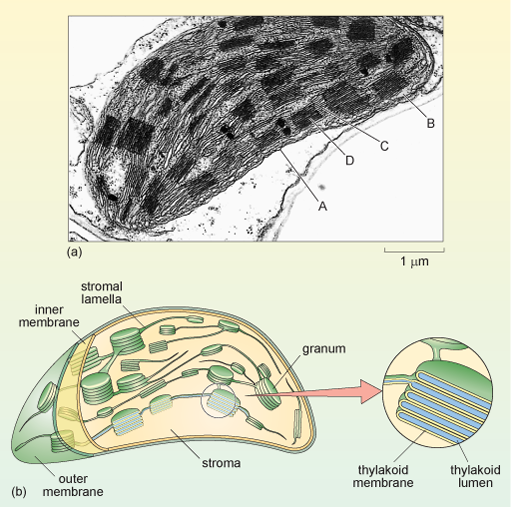
The schematic diagram in (b) shows the chloroplast bounded by an outer membrane immediately beneath which is the inner membrane. This membrane encloses the stroma, within which are the cylindrical grana, connected by membranes called stroma lamellae. The enlargement shows the internal structure of a single granum: stacks of flat membranous sacs called thylakoids each surrounding a thylakoid lumen.
The chloroplast stroma is the site of many important reactions, including the conversion of carbon dioxide into sugars, the production of starch from sugars (starch granules are often seen in chloroplasts) and the synthesis of some chloroplast proteins (there are ribosomes, DNA and RNA within the stroma).
4.12 Vacuoles
An organelle that is very prominent in all plant and fungal cells (but usually not in animal cells) is the vacuole (Figure 1b). Vacuoles can be very large and they have a number of roles. In mature plant cells, the vacuole is typically a membrane-bound space that fills most of the centre of the cell and is basically a storage compartment for water, ions and small organic molecules. In combination with the cell wall, vacuoles also have a role in determining the shape of the plant cell. A cell in which the vacuole is completely full of water is said to be turgid, and this cell turgidity ensures that the plant remains rigid and upright. In dry conditions, loss of cell turgidity results in plant wilting. Water molecules pass freely in and out of the vacuole, but the vacuolar membrane (the tonoplast) controls the passage of other molecules with specialised transporters that are different to those of the cell membrane. Vacuolar pH is lower than that of the cytosol, and vacuoles can contain enzymes that digest large organic molecules in a similar way to lysosomes in animal cells (lysosomes are rare in plant cells).
4.13 Organelle biogenesis and additional roles
There is not sufficient space in this course to describe all the properties of organelles in detail, so before leaving the topic, it is important to make two general points: firstly, organelles are dynamic structures; and secondly, they have additional roles to those described above.
New organelles form by the growth and division of existing organelles, processes that are independent of nuclear division. The formation of new organelles requires the production of new phospholipids, most of which are assembled in the SER. The new phospholipids are then incorporated into the different organelles in two ways. The first is by fusion of vesicles derived from the ER; the Golgi and lysosomes form in this way. The second is by direct incorporation of phospholipids into the organelle membrane via special membrane proteins; the mitochondria, chloroplasts and peroxisomes incorporate new phospholipids in this way, as they do not fuse with vesicles.
In addition to their main roles described in this course, many organelles are also involved in other functions. One such function is regulated cell death, often referred to as programmed cell death, one form of which is known as apoptosis. Programmed cell death is of importance in ensuring appropriate cell numbers, particularly during the development of animals and in disease. The cell membrane, mitochondria and the ER are all involved in programmed cell death.
Another function of the ER and the mitochondria is in the regulation of the levels of calcium in the cell. Calcium plays an important role in cell signalling.
Summary of Section 4
- Eukaryotic cells are surrounded by a cell membrane, which is composed of a phospholipid bilayer in which proteins and glycoproteins are embedded. The membrane forms a barrier between the cell and its external environment; membrane proteins and glycoproteins mediate selective transport of substances into and out of the cell, and interactions with the extracellular environment, including neighbouring cells.
- The eukaryotic cytoskeleton is composed of three types of filament-like structures assembled from cytoskeletal proteins: microfilaments (also known as actin filaments), microtubules and intermediate filaments. The cytoskeleton collectively provides shape, support and an internal transport system, and is responsible for cell movements.
- The nucleus, which has a double membrane (or nuclear envelope), contains most, but not all, of a cell's DNA, which is packaged by binding to histone proteins to form chromatin. DNA replication and also transcription to synthesise RNA take place in the nucleus, as does the assembly of ribosomes. RNA and assembled ribosome components leave the nucleus, and structural and ribosomal proteins and the enzymes needed for DNA replication and transcription enter it, through pores in the nuclear envelope.
- Messenger RNA is translated by ribosomes to synthesise proteins. Cytosolic proteins and proteins destined for the nucleus, mitochondria, chloroplasts and peroxisomes are synthesised by ribosomes that are free in the cytosol. Proteins destined for these organelles carry amino acid signal sequences that target them for recognition by special proteins that transport them to the appropriate cell compartments.
- Proteins destined for lysosomes, for secretion from the cell, or for embedding in the cell membrane have signal sequences that relocate the ribosome on which they are being translated to the rough ER, where their translation is completed. Glycosylation of proteins also starts in the RER.
- The smooth ER is the site of phospholipid assembly and also detoxification (for example of drugs or toxins).
- Proteins and lipids pass from the RER to the Golgi apparatus for further processing (e.g. glycosylation), and sorting and packaging into vesicles, which deliver proteins and lipids to lysosomes or to the cell membrane (either for embedding in the cell membrane, or secretion from the cell).
- Vesicles move short distances within the cell by diffusion, but are transported longer distances by movement along microtubules. Movement of vesicles to specific sites in the cell is achieved because the vesicles have special coats and surface signals that identify their contents and ensure they fuse with the appropriate membrane.
- Substances are secreted from cells by exocytosis in which vesicles fuse with the cell membrane. One way in which substances are imported into cells is by engulfment into a cell membrane-bound vesicle. This process is known as endocytosis.
- Ingested materials and 'old' organelles are digested within lysosomes. Some fatty acids and amino acids are broken down in peroxisomes.
- Mitochondria are the site of the majority of ATP production. They have a double membrane, the inner membrane of which is folded; and they also have their own DNA and ribosomes which synthesise some of the proteins required for ATP synthesis. Most mitochondrial proteins are, however, encoded by the nuclear DNA and are imported into the mitochondria.
- Plant cells have a cell wall and two organelles not found in animal cells: chloroplasts, which are the site of photosynthesis, and the vacuole. Chloroplasts have three membrane systems: outer, inner and internal thylakoid membranes. The complex internal membrane system is where photosynthesis takes place. Many other activities including synthetic reactions take place in the chloroplast stroma. The vacuole is the site where water, ions and small organic molecules are stored. It provides turgidity to the cell and may contain enzymes that digest large organic molecules.
- New organelles form by growth of pre-existing organelles, followed by division. These processes are independent of nuclear division.
- Organelles have a number of other functions, including roles in programmed cell death and calcium signalling.
5 Cell interactions in tissues
You may know from previous studies that in multicellular organisms, cells tend to be organised into groups, or tissues. This section considers the structural components that hold the constituent cells of tissues together. These are cell junctions, and an important component of all tissues, the extracellular matrix (which so far have only been mentioned in passing). The focus here is on animal tissues.
5.1 Extracellular matrix
The extracellular matrix is composed of proteins and polysaccharides that are secreted by cells and assembled into an organised meshwork external to the cells. The amount and type of extracellular matrix varies from tissue to tissue, and different types of extracellular matrix perform different functions. In many tissues, it acts rather like cement between cells; in others, it forms a support or scaffold. The extracellular matrix is not merely a scaffold, however, but can have an active role in regulating cell behaviour, and the communication between cells. Connective tissues have a relatively large proportion of extracellular material which determines their properties: for example, the rigidity of bone is due to an extracellular matrix composed of tough collagen fibres that is hardened by mineral salts (see below).
In most animal tissues, the extracellular matrix is secreted mainly by cells called fibroblasts, and the major component is a gel-like substance in which proteins are embedded. The gel is composed of polysaccharides called glycosaminoglycans (GAGs), which are often attached to proteins to form heavily glycosylated proteins called proteoglycans. The GAGs are negatively charged, and attract positively charged ions, particularly sodium ions. The high concentration of positive ions leads to the movement of water by diffusion into the matrix, giving it its gel-like properties, including the ability to resist compression.
Fibrous proteins and other substances within the extracellular matrix provide additional strength or elasticity. The most abundant extracellular protein in animal tissues is collagen, of which there are several types. Collagen is a very tough fibrous protein that can, in its different forms, provide great strength and/or flexibility (Figure 24). Elastin, as its name suggests, is a flexible protein that is found in the extracellular matrix of tissues such as blood vessels. In bone, mineral salts are laid down in connective tissue to provide support.
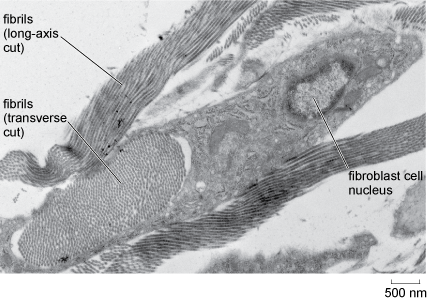
This electron micrograph shows collagen fibrils that have been sectioned along their long axis and therefore appear as linear strands orientated in the same direction. Also shown is a bundle of fibrils that have been sectioned transversely (at right-angles to their long axis). Here the fibrils have the appearance of closely spaced dark spots of uniform size. The bundle of fibrils is approximately 3 micrometres by 1 micrometre, and each fibril is estimated to be about 30 nanometres in diameter. Adjacent to and partly surrounding the fibril bundle is a fibroblast cell, which measures about 4 micrometres by 1.5 micrometres. The nucleus is darkly staining near its perimeter and the nucleolus appears as a large pale area in the centre.
Some cells are linked to particular proteins embedded in the extracellular matrix. For example, an extracellular matrix protein known as fibronectin binds to cell membrane glycoproteins known as integrins. On the inside of the cell, the integrins are linked to actin filaments of the cytoskeleton, as shown in Figure 25. Integrins can transduce information from the extracellular matrix to the cytoskeleton, which may then stimulate the cell to change shape, move or respond in other ways, to certain changes in its external environment.
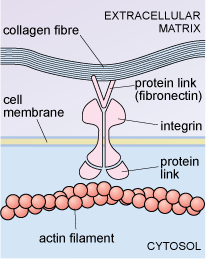
The diagram shows a membrane-spanning integrin molecule, the extracellular part of which is bound to a fibronectin molecule in the extracellular matrix. The fibronectin is also bound to a collagen molecule, and thus acts as molecular bridge between the integrin and the collagen. The cytosolic part of the integrin molecule is similarly bound via a protein linker molecule to an actin filament of the cytoskeleton.
Interactions between cells and the extracellular matrix, then, play important roles in maintaining tissue structure, and also in cell properties such as adhesion and migration.
5.2 Cell junctions
Cell-cell contacts also play extremely important roles in tissue function. For example in some tissues it is essential that there is very close contact between cells so that an effective barrier is set up and pathogenic microbes, and even harmful or unwanted molecules, cannot readily pass between the cells.
-
Give an example of circumstances where close contact between adjacent cells is important.
-
One example is in the epithelial tissue that lines the gut wall, where it is important that there are no large spaces between the cells, where harmful microbes in ingested food could enter the body. Another example is the skin, which also has a protective role.
Cells may have a number of different types of specialised cell-cell contacts known as cell junctions. Epithelial tissue, which is composed of closely packed sheets of cells has several types of cell junction each with specific functions and formed by different specialised proteins. Examples of the arrangement of the main types of junction between epithelial cells are shown in Figure 26.
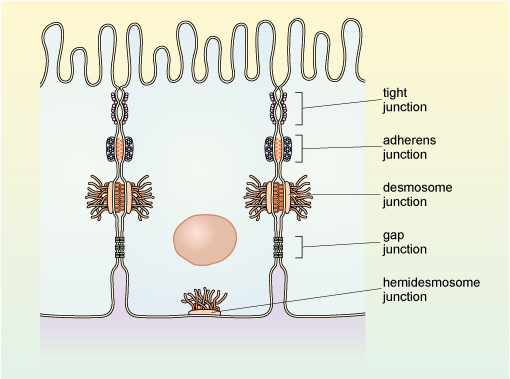
This diagram shows a small section of epithelial tissue and the location and schematic structure of the different junction types. The tight junction holds the cells very close together via their lateral surfaces, and is situated very near the apical end of the cells. The adherens junction is present on the lateral surface further from the apical end, and links the actin filaments of adjacent cells together via membrane-spanning proteins and intracellular proteins that link these to the cytoskeleton. The gap junction, which is also on the lateral surface, holds the cells tightly together: adjacent cells are separated by a narrow gap which is spanned by channel-forming protein. Also shown are the two types of anchoring junction; both of these comprise membrane-spanning adhesion proteins, which link the cells together across the intercellular space, and are linked on their cytosolic side to intermediate filaments of the cytoskeleton. The desmosome links adjacent epithelial cells together via their lateral surfaces; and the hemidesmosome links an epithelial cell to the extracellular matrix via its basal surface.
Gap junctions are a type of specialised cell junction that allows intercellular communication. In the smooth muscle of the intestine, which must contract in a coordinated manner for peristalsis to occur, the smooth muscle cells have gap junctions, which allow transmission of molecules and electrical signals between the adjacent smooth muscle cells.
Tight junctions, as their name suggests, link cells very closely forming a barrier that prevents the passage of molecules and ions between cells. They also prevent the movement of integral membrane proteins within the cell membrane. Hence they are important in maintaining, for example, the polarity of the epithelial cells of the intestine (Figure 20).
Adherens junctions link the actin cytoskeleton of adjacent cells together, often forming a belt-like arrangement around each of the cells in an epithelial sheet. The link occurs via transmembrane proteins known as cadherins, and intracellular proteins that link cadherins to the cytoskeleton.
Anchoring junctions (called desmosomes) link adjacent cells together, while hemidesmosomes link cells to the extracellular matrix. Again, these links occur via cadherins. In this case, however, the link is to intermediate filaments of the cytoskeleton rather than actin filaments.
There are other specialised types of connections between animal cells, such as the contacts between neurons and their targets, known as synapses. In plants, there are the plasmodesmata. These channels, lined by a cell membrane, make a connection between the cytoplasm of plant cells (through a gap in the cell walls), enabling transport and communication between the cells.
Summary of Section 5
- The cells of animal tissues are held together by the extracellular matrix and by cell junctions.
- The extracellular matrix in animals is composed of a hydrated mixture of glycosaminoglycans (GAGs) and proteoglycans, in which various proteins, especially fibrous proteins like collagen, are embedded. The nature of the proteins determines the properties of the matrix, which vary from tissue to tissue.
- Cells can be linked by several types of cell junctions, formed by specialised membrane proteins.
- Epithelial cells are characterised, in part, by their very close packing. They are held together by specialised junctions between adjacent cells.
Conclusion
In this course you have learnt about the subcellular components of cells and how they are studied. Cells in multicellular organisms, particularly in animals, are specialised to perform different functions; the properties, abundance and distribution of the cytoskeleton and organelles allow, and indeed underpin, this functional and structural diversity. Your study of this course has provided you with a general understanding of cellular architecture and its relationship to different processes in the cell. This sets the scene for further studies in cell biology.
Glossary
- actin
- Protein component of microfilaments of the cytoskeleton. Also found as part of the contractile apparatus of muscles.
- adherens junctions
- Junction between adjacent animal cells whose actin filaments are linked, via transmembrane proteins (cadherins) and intracellular proteins.
- anchoring junctions
- (Called desmosomes and hemidesmosomes) Intercellular junctions that are present in many types of animal tissues and serve to hold the constituent cells to each other and to the surrounding extracellular matrix respectively, via transmembrane proteins, cadherins. The link is to intermediate filaments.
- autophagy
- The degradation and recycling of the components of a cell's organelles and structures by the action of lysosomes. This process occurs when cell components are old or damaged, or in order to release nutrients under conditions of cell starvation.
- capping protein
- A protein that attaches to and stabilises microtubules and prevents disassembly.
- centrosome
- The microtubule organising centre (MTOC) found in animal cells that is involved in assembly and maintenance of the cytoskeleton. During cell division, the centrosome divides to organise the two poles of the mitotic spindle.
- cell fractionation
- Technique for the separation of the components of cells, by disruption followed by ultracentrifugation.
- chromatin
- Complex of DNA and histone proteins, which makes up eukaryotic chromosomes.
- collagen
- A protein that is abundant in the extracellular matrix. There are several types of collagen; it can form long thin fibres and provide structure, strength and flexibility to many tissues.
- cotranslational localisation
- The process whereby a ribosome with attached mRNA is moved from the cytosol to the endoplasmic reticulum (if the protein that is being encoded is destined for lysosomes, or to be secreted from the cell, or to be embedded in a cell membrane).
- cristae
- (Sing. crista) Folds of the inner mitochondrial membrane where electron transport takes place.
- cytoskeleton
- System of long filament-like protein polymers, which confer shape and support and mediate many types of cellular movement. The three types of cytoskeletal filaments are the microfilaments, microtubules and intermediate filaments.
- elastin
- A flexible protein that is found in the extracellular matrix of tissues such as blood vessels.
- endospores
- Dormant structures made by some bacteria within their cells, which can later germinate to new individual bacteria.
- euchromatin
- Regions of chromatin where the DNA is less condensed and appear pale after staining; thought to contain many genes that are being actively transcribed.
- exocytosis
- The release of materials, which may be dissolved or particulate and are enclosed within a vesicle, from a cell. Vesicles are transported to the cell surface where the cell membrane and vesicle membrane fuse, releasing the vesicle contents to the outside.
- gap junctions
- A type of specialised cell junction that allows intercellular communication, by passage of molecules and electrical signals between cells.
- glycosylation
- A type of post-translational protein modification where sugar residues are added to proteins to produce glycoproteins.
- Golgi apparatus
- A group of disc-like sacs found in eukaryotic cells, within which polysaccharides and proteins are processed (e.g. to form glycoproteins) and packaged for delivery to the cell surface for secretion to the outside.
- grana
- (Sing. granum) Stacks of thylakoids within the chloroplast. The grana are linked by stroma lamellae, which join the stacks of grana together to form a single compartment.
- granum
- See grana.
- heterochromatin
- Regions of chromatin where DNA is highly condensed, thought to be regions of chromosomes that are not transcriptionally active.
- histones
- Proteins around which DNA winds and binds to in the chromosome. The complex of DNA and histones is chromatin.
- integrins
- Type of cell surface proteins that are involved in binding to extracellular matrix components such as fibronectin, which links to collagen. On the inside of the cell, integrins link, via other proteins, to actin filaments.
- intermediate filaments
- Type of cytoskeletal filament; there are several different types of intermediate filament protein. All intermediate filaments are about 8-10 nm in diameter, and they play a role in cell support.
- intermembrane space
- The space between the inner and outer membranes in mitochondria, into which protons are pumped as a result of the activities of the mitochondrial electron transport chain (located in the inner mitochondrial membrane).
- lamins
- The type of intermediate filament found inside the nucleus.
- lysate
- The name given to the mixture of components that results when a cell/cells are disrupted, e.g by lysis (being lysed). Lysis is the term given to the rupture of the cell membrane and release of the contents of the cell.
- lysosomes
- Small spherical organelles with a single membrane. Lysosomes contain degradative enzymes and break down old organelles and also ingested material. They have a low internal pH.
- microfilaments
- Cytoskeletal proteins composed of sub-units of the protein actin. They are the thinnest of the cytoskeletal filaments, having a diameter of about 6nm, and mediate many cell movements.
- microtubules
- Tubular cytoskeletal structures composed of subunits of the protein tubulin. They are the largest of the cytoskeletal filaments, having a diameter of about 25 nm. Microtubules play an important role in maintaining cell shape and mediate the transport of vesicles and other cellular components.
- mitochondrial DNA
- The DNA that is localised in mitochondria (in the mitochondrialmatrix). It encodes most of the proteins of the inner mitochondrial membrane.
- mitochondrial matrix
- The innermost part of the mitochondrion, surrounded by a double membrane. It contains mitochondrial DNA, and the TCA cycle takes place in the mitochondrial matrix.
- nuclear pores
- Small gaps in the nuclear envelope that form channels of about 9 nm in diameter, allowing the free passage of small water-soluble molecules into and out of the nucleus. Movement of specific proteins into the nucleus, and RNA out of the nucleus is regulated by proteins of the nuclear pore complex.
- nucleoli
- See nucleolus.
- nucleolus
- Electron-dense structures in the nucleus that are the site of ribosomal RNA transcription and ribosome assembly.
- peptidoglycan
- Major component of bacterial cell walls. It is a polymer of high molecular mass, composed of two complex monosaccharides derived from glucose, which are linked together by amino acids, including three amino acids that are not found naturally in proteins.
- peroxisomes
- Small spherical organelles, bound by a single membrane. Peroxisomes contain enzymes that oxidise fatty acids and amino acids.
- phagocytosis
- The term used for engulfment of large particles or bacteria by endocytosis. Some cells of the immune system, known as phagocytes or macrophages engulf bacteria by phagocytosis.
- protein sorting
- Also called protein targeting. The delivery of newly synthesised proteins to the correct destination within the cell or, in the case of secreted proteins, to the exterior.
- protein targeting
- See protein sorting
- rough endoplasmic reticulum (RER)
- Part of the endoplasmic reticulum where protein synthesis takes place. Ribosomes are hence attached to the RER, giving it a 'rough' appearance. It is the site of synthesis of membrane and lysosomal proteins, and also proteins that are to be exported from the cell.
- scanning electron microscopy (SEM)
- A type of microscopy used to study the surface of intact cells and tissues. A sample is coated with a thin metallic layer that deflects an electron beam onto a detector.
- signal recognition particle (SRP)
- A protein complex that recognises a specific sequence of translated amino acids emerging from a free ribosome. The sequence is a signal that the protein is to be translated on the membrane of the rough endoplasmic reticulum (RER). Translation is interrupted whilst the whole complex (including the ribosome, mRNA and partially translated polypeptide) is moved to the RER.
- signal sequence
- A short, N-terminal sequence at the end of a protein molecule that serves as its 'address label', directing proteins to specific targets in the cell.
- smooth endoplasmic reticulum (SER)
- Part of the endoplasmic reticulum where phospholipids and steroids are produced, and where detoxification of drugs occurs.
- thylakoids
- Internal membrane system of chloroplasts, where photosynthesis takes place.
- tight junctions
- A type of junction that forms tight contacts between cells in animal tissues such as epithelia. Tight junctions prevent the lateral movement of other membrane proteins, thus allowing different regions of the cell membrane to have different functional properties (e.g. transport characteristics).
- transmission electron microscopy (TEM)
- A type of microscopy used to study the internal organisation of cells. A beam of electrons is passed through a very thin tissue section and an image obtained by focussing them on a fluorescent screen that emits light when electrons hit it.
- transport vesicles
- Small membrane-bound sacs that bud off from one membrane, move through the cytosol and fuse with another membrane. For example, vesicles move from ER to Golgi and from Golgi to the cell membrane and to lysosomes.
- tubulin
- Protein subunit of microtubules.
- vacuole
- Membrane-bound space, prominent in all plant and fungal cells (but usually not in animal cells), which acts as a storage compartment for water, ions and small organic molecules. In mature plant cells the vacuole fills most of the centre of the cell and together with the cell wall plays a role in determining the shape of the cell.
Acknowledgements
Except for third party materials and otherwise stated (see terms and conditions), this content is made available under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 Licence.
Course image: Umberto Salvagnin in Flickr made available under Creative Commons Attribution-NonCommercial-ShareAlike 2.0 Licence.
The material acknowledged below is Proprietary, used under licence and not subject to Creative Commons licence. See terms and conditions. Grateful acknowledgement is made to the following:
Figures
Figure 2b: Dr Gopal Murti/Science Photo Library;
Figure 3: Courtesy of Heather Davies, Department of Life, Health and Chemical Sciences, Open University;
Figure 10: Dr Torsten Wittmann/Science Photo Library;
Figure 11a, 17a, 19a and 22a: Courtesy of Professor Michael Stewart, Department of Life, Health and Chemical Sciences, Open University;
Figure 12: Adapted from Purves, W.K. et al. (1998) Life, the Science of Biology, 5th edn, Sinauer Associates Inc;
Figure 14: Dr Richard Kessel and Dr Gene Shih, Visuals Unlimited/Science Photo Library;
Figure 16: Photo printed with permission from O. L. Miller, B. A. Hamkalo and C. A. Thomas Jr;
Figure 21a: CNRI/ Science Photo Library;
Figure 21b: Dr Don Fawcett/Science Photo Library;
Figure 22c: Dr Gopal Murti/Science Photo Library;
Figure 24: Courtesy of Jill Saffrey, Department of Life, Health and Chemical Sciences, Open University.
Every effort has been made to contact copyright holders. If any have been inadvertently overlooked the publishers will be pleased to make the necessary arrangements at the first opportunity.
Don't miss out:
If reading this text has inspired you to learn more, you may be interested in joining the millions of people who discover our free learning resources and qualifications by visiting The Open University - www.open.edu/ openlearn/ free-courses
Copyright © 2016 The Open University